The respiratory electron transfer chain (ETC) of mitochondria consists of a series of large membrane-bound protein complexes (Complexes I, II, III, IV) which together with a small lipid ubiquinone (UQ) and the small protein cytochrome c catalyse the transfer of electrons from NADH and succinate to O2, forming H2O (Figure 2.25). Electron flow from NADH and succinate to oxygen is coupled to proton translocation out of the matrix to the intermembrane space which establishes a proton electrochemical gradient (DµH+) across the inner membrane that is used to drive phosphorylation of ADP to form ATP by the F1FO ATP synthase (Complex V, Figure 2.25).
Fig.2.25_edit_1.png
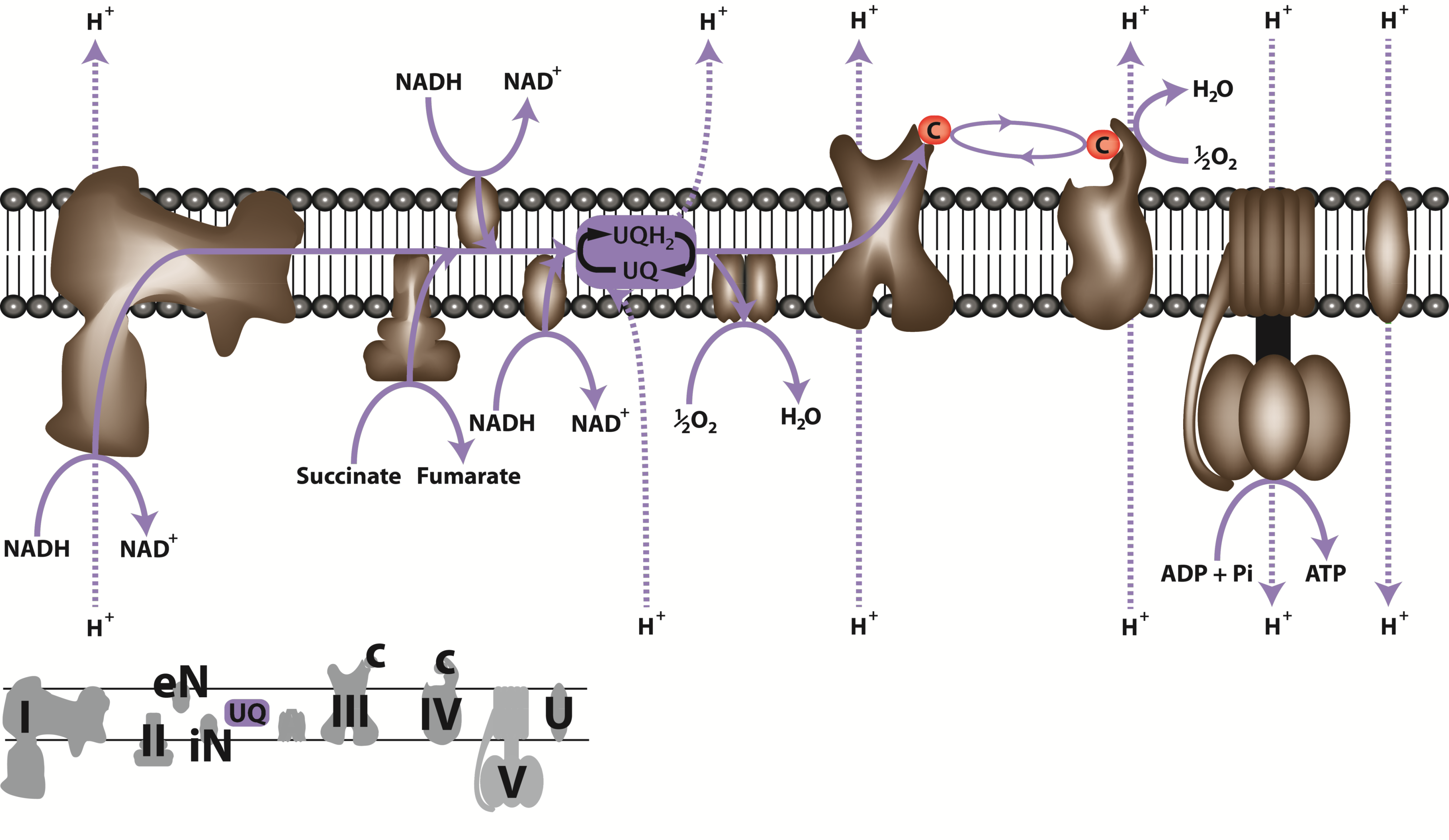
Figure 2.25. The electron transport chain of plant mitochondria. Numbers (I-V) identify the large respiratory complexes located on the inner mitochondrial membrane. Complex I, NADH:ubiquinone oxidoreductase; complex II, succinate dehydrogenase; complex III, ubiquinone-cytochrome c oxidoreductase; complex IV, cytochrome c oxidase; complex V, ATP synthase. Letters identify alternative pathway enzymes, eN, external NAD(P)H dehydrogenase; iN, internal NAD(P)H dehydrogenase; A, alternative oxidase and UQ, ubiquinone/ubiquinole pool; c, cytochrome c; U, uncoupling protein. NADH, nicotinamide adenine dinucleotide (reduced); NAD+ nicotinamide adenine dinucleotide (oxidised); ATP, adenosine triphosphate; ADP, adenosine diphosphate; Pi, inorganic phosphate. Unbroken arrows indicate pathways of electron flow; broken arrows indicate proton translocation sites. (Original drawing courtesy Nicolas Taylor & Harvey Millar)
(a) Complex I (CI)
NADH-UQ oxidoreductase, is responsible for the oxidation of matrix NADH and reduction of ubiquinone (UQ) in the inner mitochondrial membrane (Figure 2.25). In plants it is a large multi-subunit complex composed of 49 subunits, up to ten of which are synthesised in the mitochondrion whilst the others are imported from the cytosol. One of the subunits, a 50 kDa protein, contains flavin mononucleotide as a cofactor and is the dehydrogenase which oxidises NADH and passes electrons to iron-sulphur containing subunits of the complex, and eventually to ubiquinone. The passage of electrons through the complex is accompanied by H+ translocation across the membrane. Complex I is inhibited specifically by the flavonoid rotenone and its analogues. The NADH-binding site is exposed to the matrix and the complex oxidises NADH produced by the TCA cycle and other NAD-linked enzymes (Figure 2.25). Studies of mutations of CI subunits have shown that plants can survive without CI due to the activity of alternative NAD(P)H dehydrogenases (see below). Such mutants have a variety of interesting phenotypes including viral infection tolerance, prolonged hydration under water-deficient conditions and altered organic and amino acid concentrations. Using a series of Arabidopsis CI subunit knockout mutants, a number membrane arm subcomplexes (of 200, 400, 450 and 650 kDa) have been identified using BN-PAGE and antibodies. It is proposed that at least some of these subcomplexes may be assembly intermediates during CI formation, and these are seen to accumulate when specific subunits are absent.
(b) Complex II (CII)
Succinate dehydrogenase, is an enzyme of both the respiratory ETC and the TCA cycle (see above) (Figure 2.25). It is composed of four core subunits: a flavoprotein (SDHI), an iron-sulphur subunit (SDH2) and two membrane anchor subunits (SDH3 and SDH4) in most organisms. In plants, the purification of the complex has revealed the common core subunits, but also additional proteins of unknown function that co-migrate with the complex. All SDH subunits are encoded in the nuclear genome in Arabidopsis. SDH contains FeS and flavin centres which participate in electron transfer from succinate to ubiquinone. Unlike complex I, complex II does not pump H+ and succinate oxidation is therefore linked to the synthesis of less ATP per O2 reduced (see below). Malonate, an analogue of succinate, is a strong competitive inhibitor of succinate dehydrogenase. Knockout mutants of the SDH1 gene have been shown to be embryo lethal, but knockdown of SDH1 and SDH2 leads a array of phenotypes including altered stomatal aperture, mitochondrial ROS production and nitrogen use efficiency. SDHAF1 and SDHAF2 assist in CII assembly in plants and knockdown of the SDHAF2 homolog lowers SDH assembly and reduces root growth.
(c) Complex III (CIII)
Ubiquinone-cytochrome c oxidoreductase or the cytochrome b/c1 complex as it is sometimes known, contains 10 subunits including a number of bifunctional core proteins. These proteins act both in CIII function and as a matrix processing peptidase, removing targeting presequences from imported matrix synthesized proteins (Figure 2.25). A single subunit of this complex, cytochrome b, is encoded by the plant mitochondrial genome, whilst the others are encoded by the nuclear genome. It contains two b-type cytochromes, b566 and b562, cytochrome c1, an iron-sulpher protein named the Rieske iron–sulphur protein and several other polypeptides. Electron flow from ubiquinol to cytochrome c is accompanied by the translocation H+ across the membrane, via the so-called Q cycle. Various inhibitors of complex III have been discovered, with antimycin A and myxothiazol most widely used in research. The assembly of CIII is modular and includes an early core subcomplex, a late core subcomplex and the final dimeric CIII. Approximately 13 assembly factors implicated in aiding one or more of the different stages of CIII assembly in yeast, however little is known about CIII assembly or functional assembly factors in plants.
(d) Complex IV (CIV)
Cytochrome c oxidase is the final step of electron transfer of the classical ETC. As the name implies, cytochrome c oxidase accepts electrons from cytochrome c and transfers them to O2 which is reduced to form H2O. Purification of CIV in plants has identified a complex containing 14 protein subunits (Figure 2.25). Eight of these proteins are homologous to known CIV subunits from other organisms, together with a further six proteins that are probably plant specific. Two cytochrome haem centres, a and a3, and two copper atoms make up its redox active components and like complex I, cytochrome oxidase is a proton pump. Cytochrome c oxidase is sensitive to a number of inhibitors, the best known of which are carbon monoxide and cyanide. Plants, however, show resistance to both carbon monoxide and cyanide because they are equipped with an alternative oxidase (see below). Studies of human and yeast CIV has shown an assembly pathway that involves the sequential incorporation of CIV subunits, initiated by subunit 1 and assisted by over 40 assembly factors. Research investigating the plant homolog of the yeast assembly factor COX19 has found it is capable of complementing the yeast cox19 null mutant. This suggests it might play a role in the biogenesis of plant cytochrome c oxidase or in the replacement of damaged forms of the enzyme. However, our knowledge of the assembly of CIV in plants is still incomplete.
(e) Ubiquinone and Cytochrome c
These large multi-subunit complexes (I, II, III, IV) of the respiratory ETC chain are embedded in the inner mitochondrial membrane by virtue of their hydrophobic subunits, and interact with one another via two smaller molecules: ubiquinone and cytochrome c. The lipid-soluble ubiquinone also known as coenzyme Q10 is a small mobile electron carrier which moves rapidly along and across the membrane, and participates in H+ transport across the membrane via the Q-cycle as well as shuttling electrons from complexes I and II to complex III. Cytochrome c is a small haem-containing protein located on the outer surface of the inner membrane, which shuttles electrons between complexes III and IV. In this respect, the respiratory chain is similar in layout to the photosynthetic electron transport chain: three large complexes which communicate by a quinone and a small mobile protein (Cyt c or plastocyanin). However, orientation of components in the membrane is inverted and the net reaction catalysed is opposite to that in chloroplasts (Figure 1.12).
